[/vc_column_text][/vc_column][/vc_row]
This inducible resistance leads to a dissociated phenotype that is resistant to inducers (erythromycin) and susceptible to non-inducers (clindamycin).
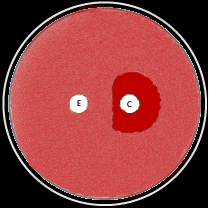
Detection of inducible resistance is important in Staphylococcus and beta-hemolytic streptococci. This is achieved by placing an erythromycin disk near to a clindamycin disk on a MH agar plate. (11,12) A D-shaped zone around the clindamycin disk (D-test positive) reveals the presence of this type of resistance mechanism (Figure 2). (9)
Figure 2. Schematic representation D-test. Organisms with inducible methylase expression are phenotypically resistant to erythromycin (inducer) but may appear susceptible to non-inducers (clindamycin). The D-test allows for the detection of inducible resistance by the modification of the inhibition zone around the clindamycin disk © when tested close to an erythromycin disk (E).
Isolates with inducible resistance to macrolides should also be reported resistant to clindamycin by the laboratory.(13)
In some strains, expression of methylase enzymes occurs in the absence of an inducer (constitutive expression) due to deletions or mutations in the erm gene conferring resistance to macrolides and clindamycin.
Enzymatic drug modification
Several enzymes produced by bacteria have been identified to have the ability to modify these antibiotics.
Esterases and phosphotransferases produced by Enterobacteriaceae inactivate the lactone ring of macrolides conferring resistance.
Phosphotransferases and nucleotidyltransferases have been associated with resistance to lincosamides in Streptomyces, Staphylococcus, Bacteroides, and Streptococcus agalactiae. (5,6)
Efflux
Active efflux of antibiotics by the expression of efflux pumps has been linked to resistance in gram positive organisms and it is responsible for the intrinsic resistance to macrolides and lincosamides of Escherichia coli and Enterococcus faecalis. (14)



